Teaching Machines to Read Radiology Reports
At Qure, we build deep learning models to detect abnormalities from radiological images. These models require huge amount of labeled data to learn to diagnose abnormalities from the scans. So, we collected a large dataset from several centers, which included both in-hospital and outpatient radiology centers. These datasets contain scans and the associated clinical radiology reports.
For now, we use radiologist reports as the gold standard as we train deep learning algorithms to recognize abnormalities on radiology images. While this is not ideal for many reasons (see this), it is currently the most scalable way to supply classification algorithms with the millions of images that they need in order to achieve high accuracy.
These reports are usually written in free form text rather than in a structured format. So, we have designed a rule based Natural Language Processing (NLP) system to extract findings automatically from these unstructured reports.
CT SCAN BRAIN - PLAIN STUDY Axial ct sections of the brain were performed from the level of base of skull. 5mm sections were done for the posterior fossa and 5 mm sections for the supra sellar region without contrast. OBSERVATIONS: - Area of intracerebral haemorrhage measuring 16x15mm seen in left gangliocapsular region and left corona radiate. - Minimal squashing of left lateral ventricle noted without any appreciable midline shift - Lacunar infarcts seen in both gangliocapsular regions - Cerebellar parenchyma is normal. - Fourth ventricle is normal in position and caliber. - The cerebellopontine cisterns, basal cisterns and sylvian cisterns appear normal. - Midbrain and pontine structures are normal. - Sella and para sellar regions appear normal. - The grey-white matter attenuation pattern is normal. - Calvarium appears normal - Ethmoid and right maxillary sinusitis noted IMPRESSION: - INTRACEREBRAL HAEMORRHAGE IN LEFT GANGLIOCAPSULAR REGION AND LEFT CORONA RADIATA - LACUNAR INFARCTS IN BOTH GANGLIOCAPSULAR REGIONS
⇩
{
"intracerebral hemorrhage": true,
"lacunar infarct": true,
"mass effect": true,
"midline shift": false,
"maxillary sinusitis": true
}
An example clinical radiology report and the automatically extracted findings
Why Rule based NLP ?
Rule based NLP systems use a list of manually created rules to parse the unorganized content and structure it. Machine Learning (ML) based NLP systems, on the other hand, automatically generate the rules when trained on a large annotated dataset.
Rule based approaches have multiple advantages when compared to ML based ones:
- Clinical knowledge can be manually incorporated into a rule based system. Whereas, to capture this knowledge in a ML based system, a huge amount of annotation is required.
- Auto-generated rules of ML systems are difficult to interpret compared to the manually curated rules.
- Rules can be readily added or modified to accommodate a new set of target findings in a rule based system.
- Previous works on clinical report parsing[1, 2] show that the results of machine learning based NLP systems are inferior to that of rule based ones.
Development of Rule based NLP
As reports were collected from multiple centers, there were multiple reporting standards. Therefore, we constructed a set of rules to capture these variations after manually reading a large number of reports. Of these, I illustrate two common types of rules below.
Findings Detection
In reports, the same finding can be noted in several different formats. These include the definition of the finding itself or its synonyms. For example, finding blunted CP angle could be reported in either of the following ways:
- CP angle is obliterated
- Hazy costophrenic angles
- Obscured CP angle
- Effusion/thickening
We collected all the wordings that can be used to report findings and created a rule for each finding. As an illustration, following is the rule for blunted CP angle.
((angle & (blunt | obscur | oblitera | haz | opaci)) | (effusio & thicken))
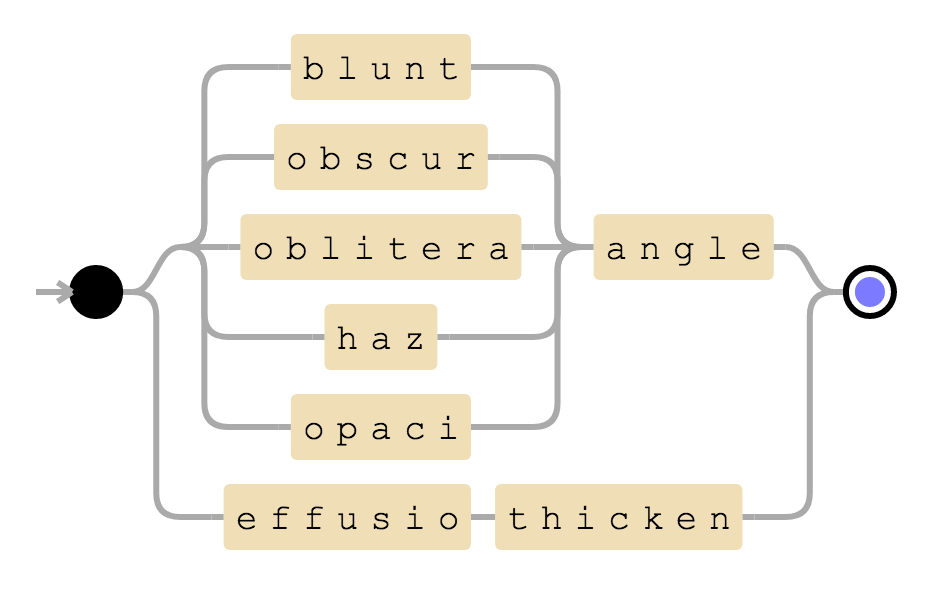
Visualization of blunted CP angle rule
This rule will be positive if there are words angle and blunted or its synonyms in a sentence. Alternatively, it will also be positive if there are words effusion and thickening in a given sentence.
In addition, there can be a hierarchical structure in findings. For example, opacity is considered positive if any of the edema, groundglass, consolidation etc are positive. We therefore created a ontology of findings and rules to deal with this hierarchy.
[opacity] rule = ((opacit & !(/ & collapse)) | infiltrate | hyperdensit) hierarchy = (edema | groundglass | consolidation | ... )
Rule and hierarchy for opacity
Negation Detection
The above mentioned rules are used to detect a finding in a report. But these are not sufficient to understand the reports. For example, consider the following sentences.
1. Intracerebral hemorrhage is absent. 2. Contusions are ruled out. 3. No evidence of intracranial hemorrhages in the brain.
Although the findings intracerebral hemorrhage, contusion and intracranial hemorrhage are mentioned in the above sentences, their absence is noted in these sentences rather than their presence. Therefore, we need to detect negations in a sentence in addition to findings.
We manually read several sentences that indicate negation of findings and grouped these sentences according to their structures. Rules to detect negation were created based on these groups. One of these is illustrated below:
(<finding>) & ( is | are | was | were ) & (absent | ruled out | unlikely | negative)
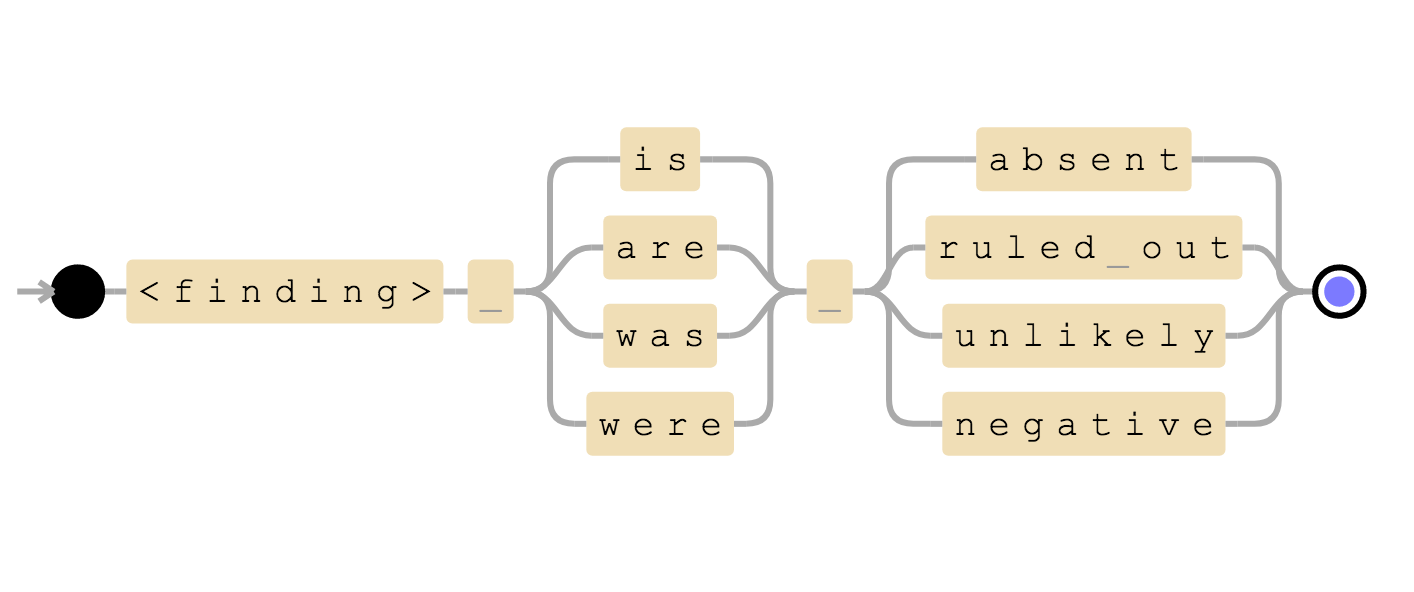
Negation detection structure
We can see that first and second sentences of above example matches this rule and therefore we can infer that the finding is negative.
- Intracerebral hemorrhage is absent ⟶
intracerebral hemorrhagenegative. - Contusions are ruled out ⟶
contusionnegative.
Results:
We have tested our algorithm on a dataset containing 1878 clinical radiology reports of Head CT scans. We manually read all the reports to create gold standards. We used sensitivity and specificity as evaluation metrics. The results obtained are given below in a table.
| Findings | #Positives | Sensitivity (95% CI) | Specificity (95% CI) |
|---|---|---|---|
| Intracranial Hemorrhage | 207 | 0.9807 (0.9513-0.9947) | 0.9873 (0.9804-0.9922) |
| Intraparenchymal Hemorrhage | 157 | 0.9809 (0.9452-0.9960) | 0.9883 (0.9818-0.9929) |
| Intraventricular Hemorrhage | 44 | 1.0000 (0.9196-1.0000) | 1.0000 (0.9979-1.0000) |
| Subdural Hemorrhage | 44 | 0.9318 (0.8134-0.9857) | 0.9965 (0.9925-0.9987) |
| Extradural Hemorrhage | 27 | 1.0000 (0.8723-1.0000) | 0.9983 (0.9950-0.9996) |
| Subarachnoid Hemorrhage | 51 | 1.0000 (0.9302-1.0000) | 0.9971 (0.9933-0.9991) |
| Fracture | 143 | 1.0000 (0.9745-1.0000) | 1.0000 (0.9977-1.0000) |
| Calvarial Fracture | 89 | 0.9888 (0.9390-0.9997) | 0.9947 (0.9899-0.9976) |
| Midline Shift | 54 | 0.9815 (0.9011-0.9995) | 1.0000 (0.9979-1.0000) |
| Mass Effect | 132 | 0.9773 (0.9350-0.9953) | 0.9933 (0.9881-0.9967) |
In this paper[1], authors used ML based NLP model (Bag Of Words with unigrams, bigrams, and trigrams plus average word embeddings vector) to extract findings from head CT clinical radiology reports. They reported average sensitivity and average specificity of 0.9025 and 0.9172 across findings. The same across target findings on our evaluation turns out to be 0.9841 and 0.9956 respectively. So, we can conclude rule based NLP algorithms perform better than ML based NLP algorithms on clinical reports.
References
- John Zech, Margaret Pain, Joseph Titano, Marcus Badgeley, Javin Schefflein, Andres Su, Anthony Costa, Joshua Bederson, Joseph Lehar & Eric Karl Oermann (2018). Natural Language–based Machine Learning Models for the Annotation of Clinical Radiology Reports. Radiology.
- Bethany Percha, Houssam Nassif, Jafi Lipson, Elizabeth Burnside & Daniel Rubin (2012). Automatic classification of mammography reports by BI-RADS breast tissue composition class.